


Giving field researchers tools to validate their own data, against standards set by their own data management plan, ensures that all downstream users of this data can rely on accurate data when it counts. Our open source software verifies that data conforms to a known standard and checks data format, numerical ranges, controlled vocabularies, and dependency checks. Data quality tests are configurable depending on your intended downstream use of the collected data.
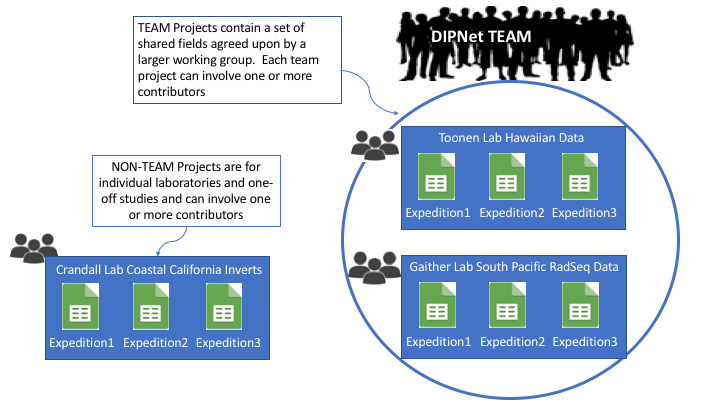
We help organizations, teams, and scientific projects to create plans for managing data, while connecting to community standard vocabularies. We enable teams to build on a global set of rules while creating an environment for individual members to interact and contribute data. Our tools are built to be adaptable to many different types of data. One example network is the Genomics Observatory Metadatabase, listed under projects below.
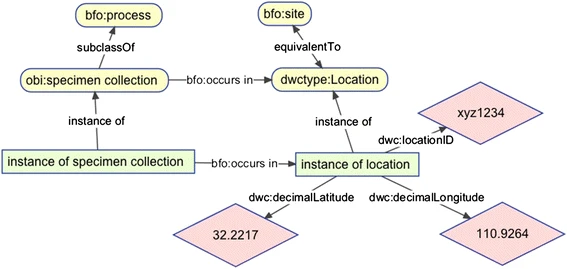
Ontologies help in data integration by using a logic-based framework to define relationships between entities. Ontologies are a powerful tool, but often users are presented with the challenge of using ontologies effectively in data management. We have built a flexible, scalable pipeline for integration and alignment of multiple data sources using ontologies. Processing is adaptable to all kinds of data or reasoning profiles, and output is compatible with any type of storage technology. The ontology-data-pipeline is designed to be run as a Docker container but can also be run natively in python. For more information visit the ontology-data-pipeline github page.
The Genomic Observatories Meta-Database (GEOME) is a web-based database that captures the who, what, where, and when of biological samples and associated genetic sequences. GEOME helps users with ensuring metadata from samples is FAIR, improving data quality, and enabling integration with downstream tools
Using the ontology-data-pipeline, the global plant phenology portal integrates data from diverse sources, assembling over 20 million phenological observations and aligning with the Plant Phenology Ontology
FuTRES (Functional Trait Resource for Environmental Studies) is a workflow for assembling functional trait data measured at the specimen level, and a database to serve that data. It is based on a semantic model and is powered by extensible parsers, a backend database, and an API.
The Biocode LIMS software operates as a plugin to the Geneious software and comprises everything you need to manage your lab and sequence analysis workflows. Our wiki serves both as a repository for the plugins themselves (with release notes and links to downloads), and an extensive online manual.
Biocode, LLC team members are listed below. If you want information about Biocode, LLC, our technology options or any of our projects, do not hesitate to reach out by sending an email to biocodellc@gmail.com
Founding member and project management
API and Data Integration